Overview
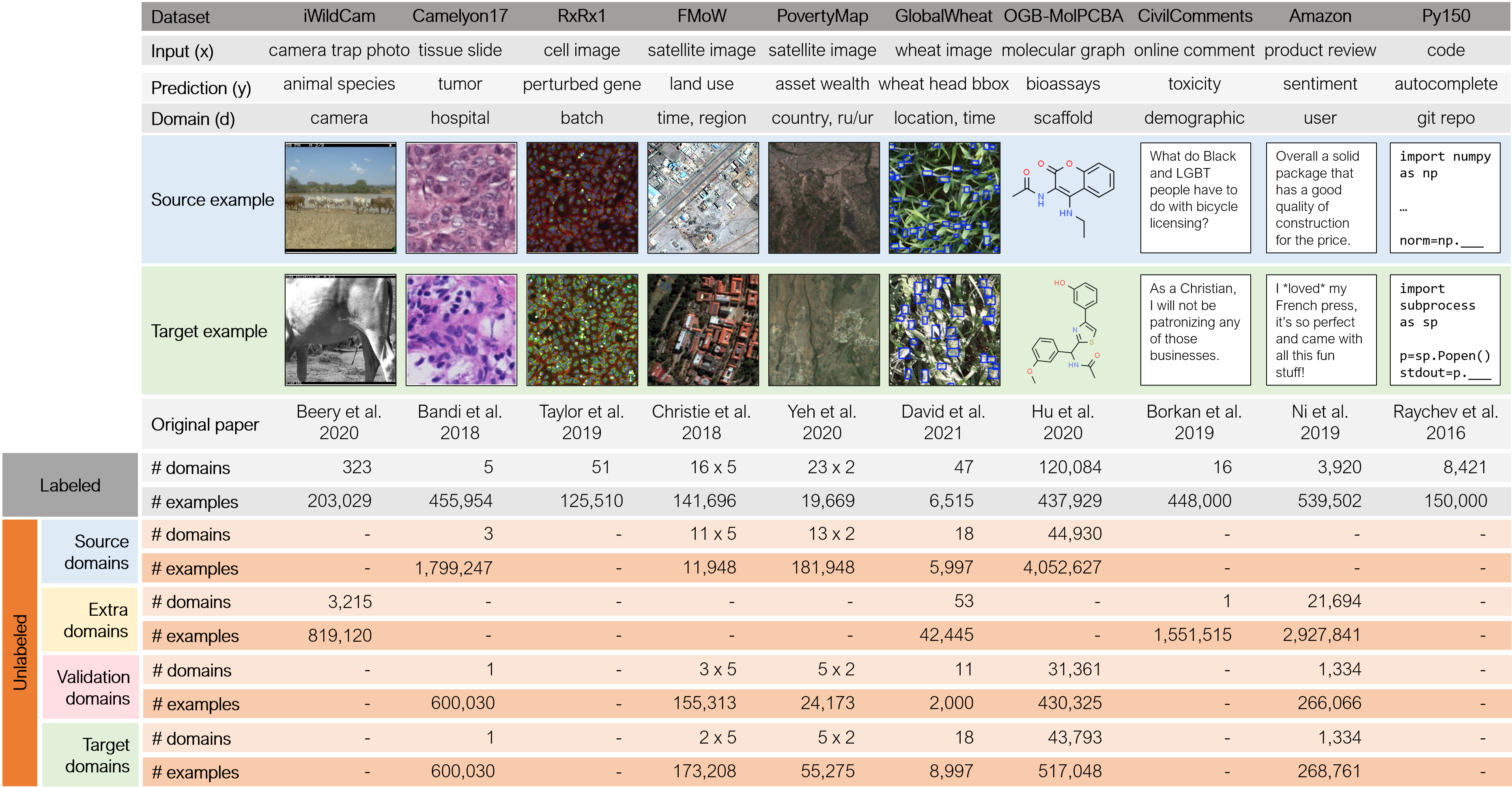
WILDS is a curated collection of benchmark datasets that represent distribution shifts faced in the wild. In each dataset, each data point is drawn from a domain, which represents a distribution over data that is similar in some way, e.g., molecules with the same scaffold structure, or satellite images from the same region. We study two types of distribution shifts over domains. In domain generalization, the training and test distributions comprise disjoint sets of domains, and the goal is to generalize to domains unseen during training, e.g., molecules with a new scaffold structure. In subpopulation shift, the training and test domains overlap, but their relative proportions differ. We typically assess models by their worst performance over test domains, each of which correspond to a subpopulation of interest, e.g., different geographical regions.
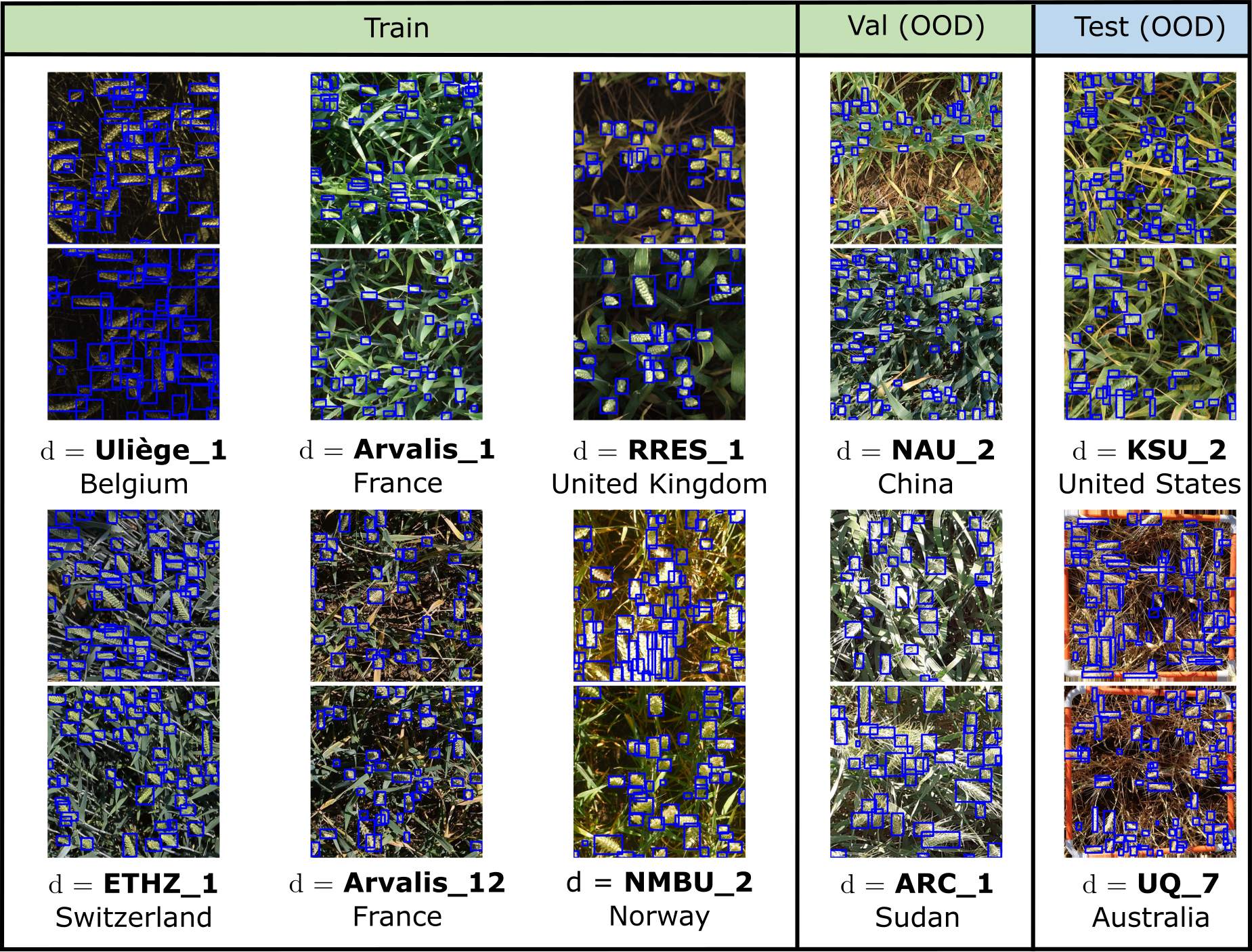
All datasets have labeled training, validation, and test data. Most datasets also have unlabeled data that can be used for training. This unlabeled data can come from the same source domains as the labeled training data; the same target domains as the labeled test data; or even extra domains that are not present in any of the labeled data. As an illustration, we show a few domains from the GlobalWheat-WILDS dataset:

Datasets
WILDS datasets span a diverse array of modalities and applications, and reflect a wide range of distribution shifts arising from different demographics, users, hospitals, camera locations, countries, time periods, and molecular scaffolds.
Click on this image to zoom in:
We are actively maintaining WILDS. To be notified of updates, please sign up for our mailing list. WILDS is a community effort, and we highly welcome contributions of real-world distribution shift datasets. Please contact us at wilds@cs.stanford.edu if you are interested in contributing.
iWildCam
Motivation
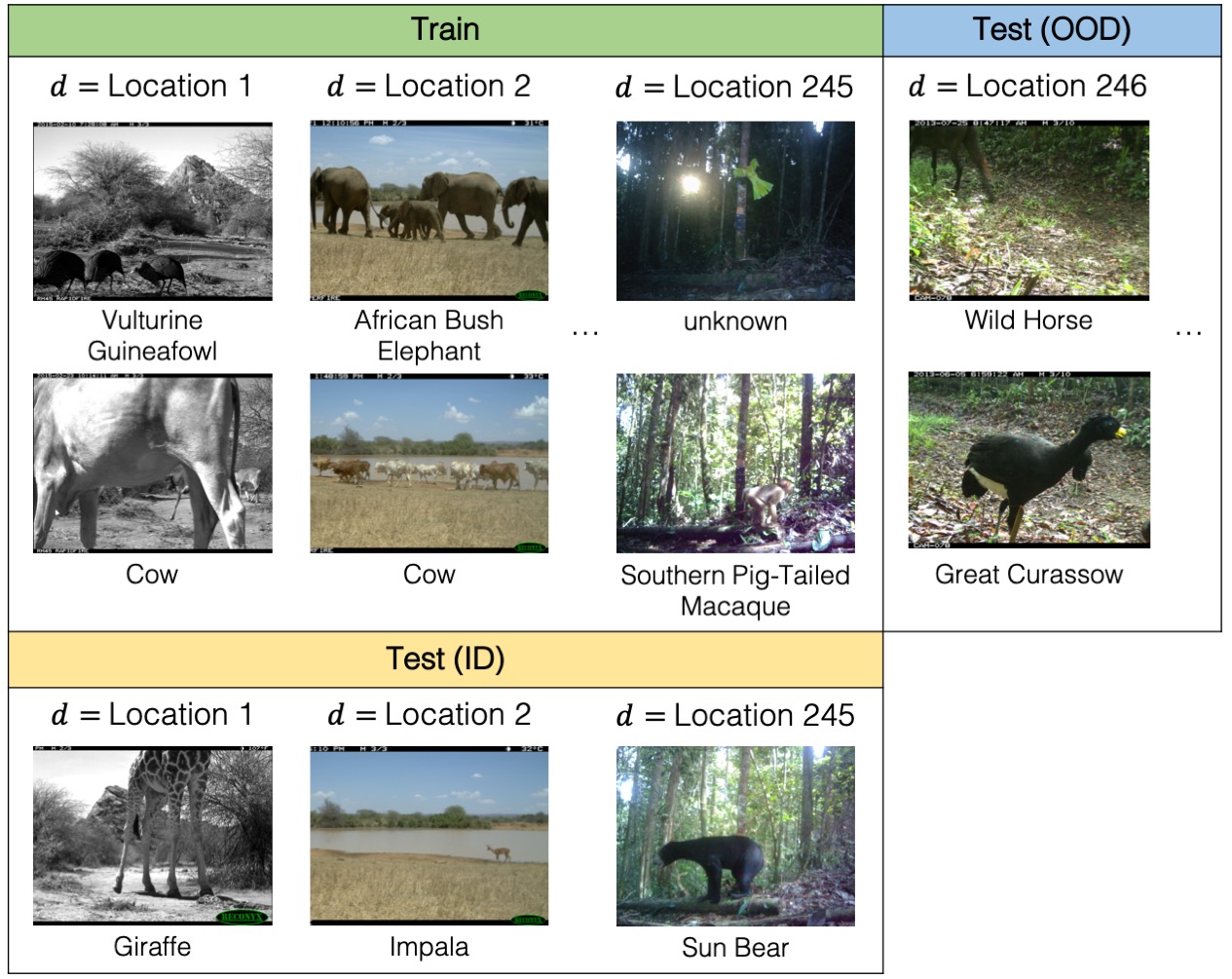
Animal populations have declined 68% on average since 1970 (Grooten et al., 2020). To better understand and monitor wildlife biodiversity loss, ecologists commonly deploy camera traps—heat or motion-activated static cameras placed in the wild (Wearn and Glover-Kapfer, 2017)—and then use ML models to process the data collected (Ahumada et al., 2020; Weinstein, 2018; Norouzzadeh et al.,2019; Tabak et al., 2019; Beery et al., 2019). Typically, these models would be trained on photos from some existing camera traps and then used across new camera trap deployments. However, across different camera traps, there is drastic variation in illumination, camera angle, background, vegetation, color, and relative animal frequencies, which results in models generalizing poorly to new camera trap deployments (Beery et al., 2018).
We study this shift on a variant of the iWildCam 2020 dataset (Beery et al., 2020).
Problem setting
We consider the domain generalization setting, where the domains are camera traps, and we seek to learn models that generalize to photos taken from new camera deployments. The task is multi-class species classification. Concretely, the input x is a photo taken by a camera trap, the label y is one of 182 different animal species, and the domain d is an integer that identifies the camera trap that took the photo.
Dataset citation
@article{beery2020iwildcam,
title={The iWildCam 2020 Competition Dataset},
author={Beery, Sara and Cole, Elijah and Gjoka, Arvi},
journal={arXiv preprint arXiv:2004.10340},
year={2020}
}
License
Distributed under the Community Data License Agreement – Permissive – V1.0.
Camelyon17
Motivation
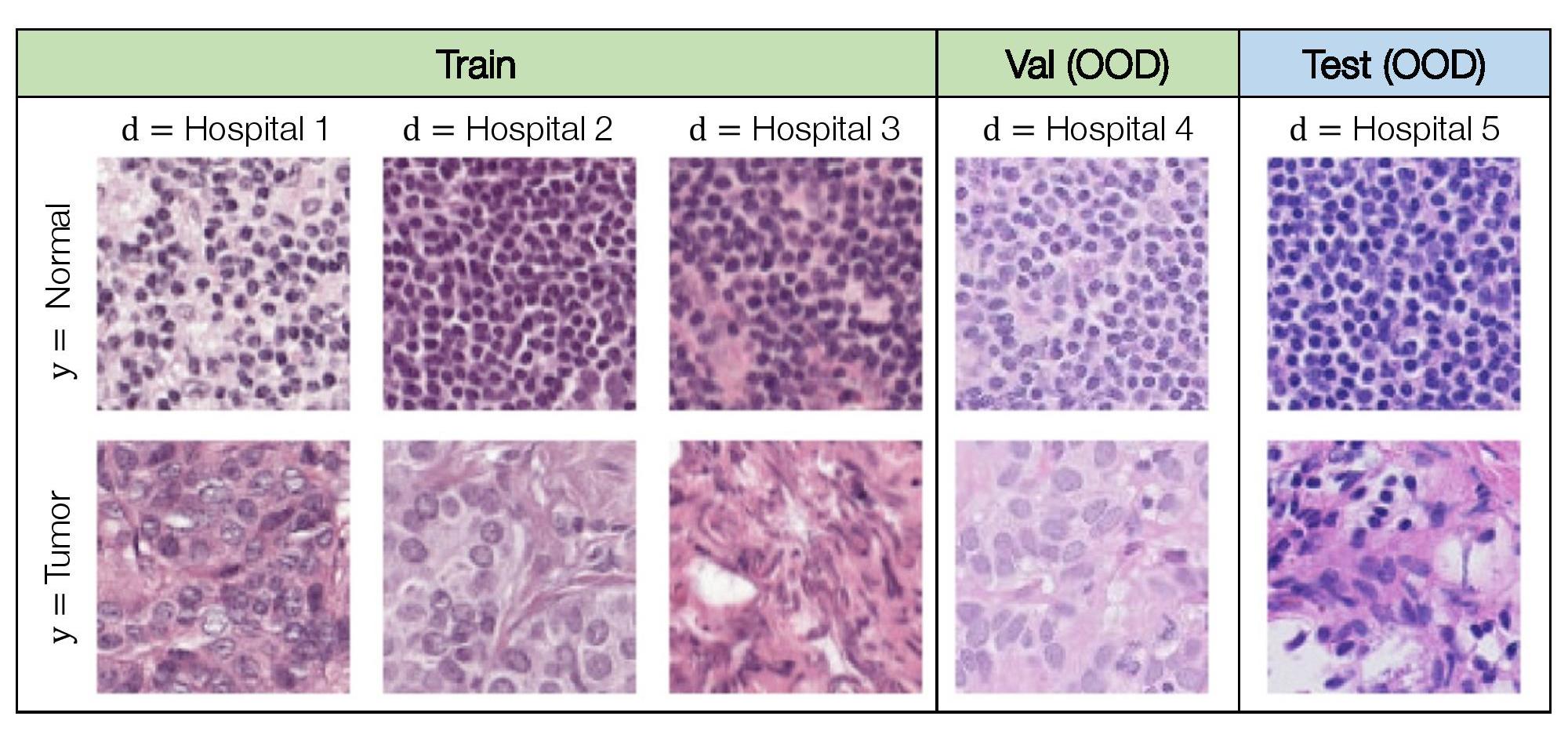
Models for medical applications are often trained on data from a small number of hospitals, but with the goal of being deployed more generally across other hospitals. However, variations in data collection and processing can degrade model accuracy on data from new hospital deployments (e.g., Zech et al. (2018); AlBadawy et al. (2018)). In histopathology applications—studying tissue slides under a microscope—this variation can arise from sources like differences in the patient population or in slide staining and image acquisition (Veta et al., 2016; Komura and Ishikawa, 2018; Tellez et al., 2019).
We study this shift on a patch-based variant of the Camelyon17 dataset (Bandi et al., 2018).
Problem setting
We consider the domain generalization setting, where the domains are hospitals, and our goal is to learn models that generalize to data from a hospital that is not in the training set. The task is to predict if a given region of tissue contains any tumor tissue, which we model as binary classification. Concretely, the input x is a 96x96 histopathological image, the label y is a binary indicator of whether the central 32x32 region contains any tumor tissue, and the domain d is an integer that identifies the hospital that the patch was taken from.
Dataset citation
@article{bandi2018detection,
title={From detection of individual metastases to classification of lymph node status at the patient level: the CAMELYON17 challenge},
author={Bandi, Peter and Geessink, Oscar and Manson, Quirine and Van Dijk, Marcory and Balkenhol, Maschenka and Hermsen, Meyke and Bejnordi, Babak Ehteshami and Lee, Byungjae and Paeng, Kyunghyun and Zhong, Aoxiao and others},
journal={IEEE Transactions on Medical Imaging},
year={2018},
publisher={IEEE}
}
License
This dataset is in the public domain and is distributed under CC0.
RxRx1
Motivation
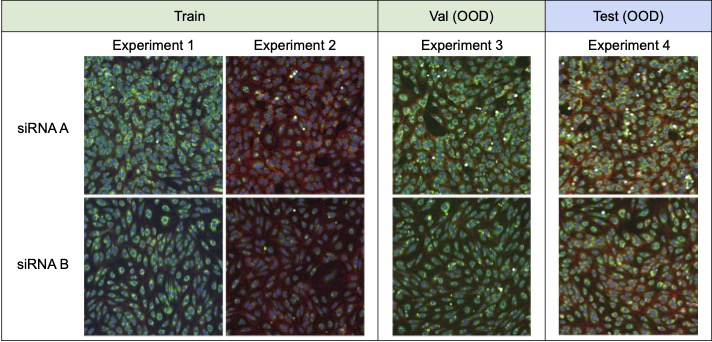
High-throughput screening techniques that can generate large amounts of data are now common in many fields of biology, including transcriptomics (Harrill et al., 2019) genomics (Echeverri and Perrimon, 2006; Zhou et al., 2014), proteomics and metabolomics (Taylor et al., 2021), and drug discovery (Broach et al., 1996; Macarron et al., 2011; Swinney and Anthony, 2011; Boutros et al., 2015). Such large volumes of data, however, need to be created in experimental batches, or groups of experiments executed at similar times under similar conditions. Despite attempts to carefully control experimental variables such as temperature, humidity, and reagent concentration, measurements from these screens are confounded by technical artifacts that arise from differences in the execution of each batch. These batch effects make it difficult to draw conclusions from data across experimental batches (Leek et al., 2010; Parker and Leek, 2012; Soneson et al., 2014; Nygaard et al., 2016; Caicedo et al., 2017).
We study the shift induced by batch effects on a variant of the RxRx1 dataset (Taylor et al., 2019).
Problem setting
We consider a domain generalization problem, where the input x is a 3-channel image of cells obtained by fluorescent microscopy (Bray et al., 2016), the label y indicates which of the 1,139 genetic treatments (including no treatment) the cells received, and the domain d specifies the batch in which the imaging experiment was run. The goal is to generalize to new experimental batches, which requires models to isolate the biological signal from batch effects.
Dataset citation
@inproceedings{taylor2019rxrx1,
author = {Taylor, J. and Earnshaw, B. and Mabey, B. and Victors, M. and Yosinski, J.},
title = {RxRx1: An Image Set for Cellular Morphological Variation Across Many Experimental Batches.},
year = {2019},
booktitle = {International Conference on Learning Representations (ICLR)},
booksubtitle = {AI for Social Good Workshop},
}
License
This dataset is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
OGB-MolPCBA
Motivation
Accurate prediction of the biochemical properties of small molecules can significantly accelerate drug discovery by reducing the need for expensive lab experiments (Shoichet, 2004; Hughes et al., 2011). However, the experimental data available for training such models is limited compared to the extremely diverse and combinatorially large universe of candidate molecules that we would want to make predictions on (Bohacek et al, 1991; Sterling and Irwin, 2015; Lyu et al, 2019; McCloskey et al, 2019). This means that models need to generalize to out-of-distribution molecules that are structurally different from those seen in the training set.
We study this issue through the OGB-MolPCBA dataset, which is directly adopted from the Open Graph Benchmark (Hu et al., 2020b) and originally curated by the MoleculeNet (Wu et al. 2018).
Problem setting
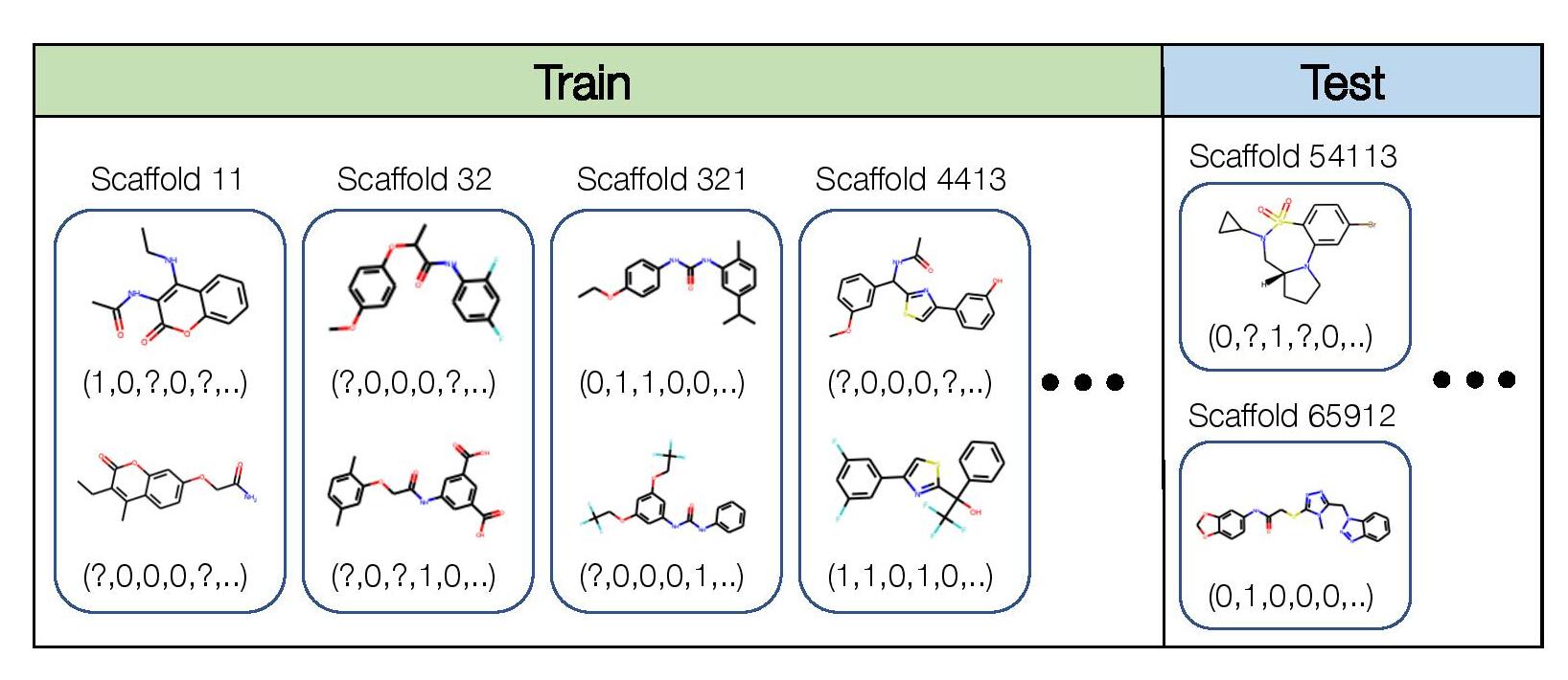
We consider the domain generalization setting, where the domains are molecular scaffolds, and our goal is to learn models that generalize to structurally-distinct molecules with scaffolds that are not in the training set. This is a multi-task classification problem: for each molecule, we predict the presence or absence of 128 different kinds of biological activities, such as binding to a particular enzyme. In addition, we cluster the molecules into different scaffold groups according to their two-dimensional structure, and annotate each molecule with the scaffold group that it belongs to. Concretely, the input x is a molecular graph, the label y is a 128-dimensional binary vector where each component corresponds to a biochemical assay result, and the domain d specifies the scaffold. Not all biological activities are measured for each molecule, so y can have missing values.
Dataset citation
@inproceedings{hu2020open,
title={Open Graph Benchmark: Datasets for machine learning on graphs},
author={Hu, Weihua and Fey, Matthias and Zitnik, Marinka and Dong, Yuxiao and Ren, Hongyu and Liu, Bowen and Catasta, Michele and Leskovec, Jure},
booktitle = {Advances in Neural Information Processing Systems (NeurIPS)},
year = {2020}
}
License
Distributed under the MIT license.
GlobalWheat
Motivation
Models for automated, high-throughput plant phenotyping—measuring the physical characteristics of plants and crops, such as wheat head density and counts—are important tools for crop breeding (Thorp et al., 2018; Reynolds et al., 2020) and agricultural field management (Shi et al., 2016). These models are typically trained on data collected in a limited number of regions, even for crops grown worldwide such as wheat (Madec et al., 2019; Xiong et al., 2019; Ubbens et al., 2020; Ayalew et al., 2020). However, there can be substantial variation between regions, due to differences in crop varieties, growing conditions, and data collection protocols. Prior work on wheat head detection has shown that this variation can significantly degrade model performance on regions unseen during training (David et al., 2020).
We study this shift in an expanded version of the Global Wheat Head Dataset (David et al., 2020; David et al., 2021), a large set of wheat images collected from 12 countries around the world.
Problem setting
We consider the domain generalization problem on a wheat head detection task, where the domains are image acquisition sessions (i.e., a specific location, time, and sensor with which a set of images was collected). The goal is to generalize to new acquisition sessions, in particular to those from regions unseen during training time; the training and test sets comprising images from disjoint continents. Concretely, the input x is a cropped overhead image of a wheat field, the label y is the set of bounding boxes for each wheat head visible in the image, and the domain d specifies an image acquisition session
Dataset citation
@article{david2020global,
title={Global Wheat Head Detection (GWHD) dataset: a large and diverse dataset of high-resolution RGB-labelled images to develop and benchmark wheat head detection methods},
author={David, Etienne and Madec, Simon and Sadeghi-Tehran, Pouria and Aasen, Helge and Zheng, Bangyou and Liu, Shouyang and Kirchgessner, Norbert and Ishikawa, Goro and Nagasawa, Koichi and Badhon, Minhajul A and Curtis Pozniak and Benoit de Solan and Andreas Hund and Scott C. Chapman and Frederic Baret and Ian Stavness and Wei Guo},
journal={Plant Phenomics},
volume={2020},
year={2020},
publisher={Science Partner Journal}
}
@misc{david2021global,
title={Global Wheat Head Dataset 2021: an update to improve the benchmarking wheat head localization with more diversity},
author={Etienne David and Mario Serouart and Daniel Smith and Simon Madec and Kaaviya Velumani and Shouyang Liu and Xu Wang and Francisco Pinto Espinosa and Shahameh Shafiee and Izzat S. A. Tahir and Hisashi Tsujimoto and Shuhei Nasuda and Bangyou Zheng and Norbert Kichgessner and Helge Aasen and Andreas Hund and Pouria Sadhegi-Tehran and Koichi Nagasawa and Goro Ishikawa and Sebastien Dandrifosse and Alexis Carlier and Benoit Mercatoris and Ken Kuroki and Haozhou Wang and Masanori Ishii and Minhajul A. Badhon and Curtis Pozniak and David Shaner LeBauer and Morten Lilimo and Jesse Poland and Scott Chapman and Benoit de Solan and Frederic Baret and Ian Stavness and Wei Guo},
year={2021},
eprint={2105.07660},
archivePrefix={arXiv},
primaryClass={cs.CV}
}
License
Distributed under the MIT license.
CivilComments

Motivation
Automatic review of user-generated text—e.g., detecting toxic comments—is an important tool for moderating the sheer volume of text written on the Internet. Unfortunately, prior work has shown that such toxicity classifiers pick up on biases in the training data and spuriously associate toxicity with the mention of certain demographics (Park et al., 2018; Dixon et al., 2018). These types of spurious correlations can significantly degrade model performance on particular subpopulations (Sagawa et al.,2020).
We study this issue through a modified variant of the CivilComments dataset (Borkan et al.,2019b).
Problem setting
We cast CivilComments as a subpopulation shift problem, where the subpopulations correspond to different demographic identities, and our goal is to do well on all subpopulations (and not just on average across these subpopulations). Specifically, we focus on mitigating biases with respect to comments that mention particular demographic identities, and not comments written by members of those demographic identities; we discuss this distinction in the broader context section below.
The task is a binary classification task of determining if a comment is toxic. Concretely, the input x is a comment on an online article (comprising one or more sentences of text) and the label y is whether it is rated toxic or not. In CivilComments, unlike in most of the other datasets we consider, the domain annotation d is a multi-dimensional binary vector, with the 8 dimensions corresponding to whether the comment mentions each of the 8 demographic identities male, female, LGBTQ, Christian, Muslim, other religions, Black, and White.
Dataset citation
@inproceedings{borkan2019nuanced,
title={Nuanced metrics for measuring unintended bias with real data for text classification},
author={Borkan, Daniel and Dixon, Lucas and Sorensen, Jeffrey and Thain, Nithum and Vasserman, Lucy},
booktitle={Companion Proceedings of The 2019 World Wide Web Conference},
year={2019}
}
License
This dataset is in the public domain and is distributed under CC0.
FMoW
Motivation
ML models for satellite imagery can enable global-scale monitoring of sustainability and economic challenges, aiding policy and humanitarian efforts in applications such as tracking deforestation (Hansen et al., 2013), population density mapping (Tiecke et al., 2017), crop yield prediction (Wang et al., 2020b), and other economic tracking applications (Katona et al., 2018). As satellite data constantly changes due to human activity and environmental processes, these models must be robust to distribution shifts over time. Moreover, as there can be disparities in the data available between regions, these models should ideally have uniformly-high accuracies instead of only doing well on data-rich regions and countries.
We study this problem on a variant of the Functional Map of the World dataset (Christie et al., 2018).
Problem setting
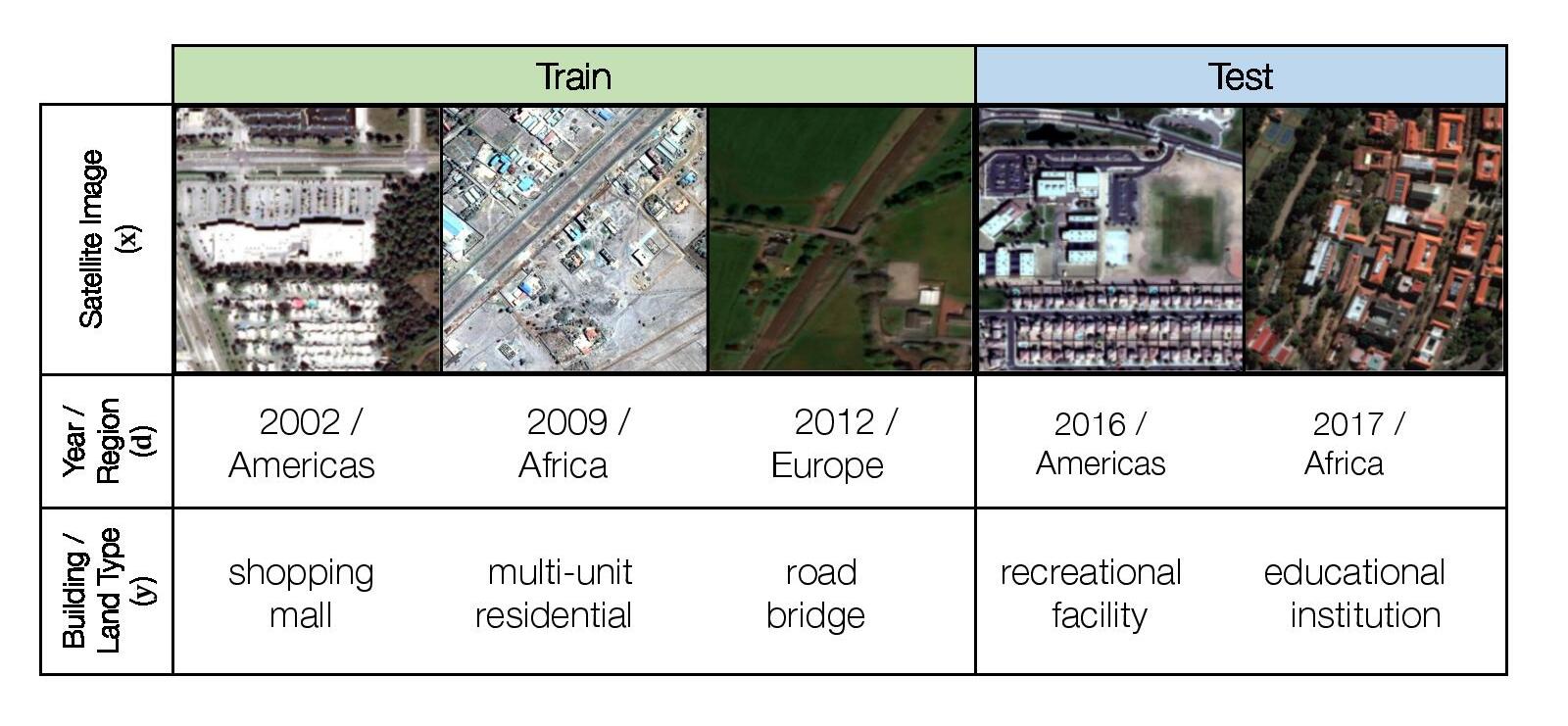
We consider a hybrid domain generalization and subpopulation shift problem, where the input x is a RGB satellite image (resized to 224 x 224 pixels), the label y is one of 62 building or land use categories, and the domain d represents both the year the image was taken as well as its geographical region (Africa, the Americas, Oceania, Asia, or Europe). We aim to solve both a domain generalization problem across time and improve subpopulation performance across regions.
Dataset citation
@inproceedings{christie2018functional,
title={Functional Map of the World},
author={Christie, Gordon and Fendley, Neil and Wilson, James and Mukherjee, Ryan},
booktitle={Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition},
year={2018}
}
License
Distributed under the FMoW Challenge Public License.
PovertyMap
Motivation
A different application of satellite imagery is poverty estimation across different spatial regions, which is essential for targeted humanitarian efforts in poor regions (Espey et al., 2015; Abelson et al., 2014). However, ground-truth measurements of poverty are lacking for much of the developing world, as field surveys are expensive (Blumenstock et al., 2015; Xie et al., 2016; Jean et al., 2016). For example, at least 4 years pass between nationally representative consumption or asset wealth surveys in the majority of African countries, with seven countries that had either never conducted a survey or had gaps of over a decade between surveys (Yeh et al., 2020). One approach to this problem is to train ML models on countries with ground truth labels and then deploy them to different coutries where we have satellite data but no labels.
We study this problem through a variant of the poverty mapping dataset collected by Yeh et al.(2020).
Problem setting
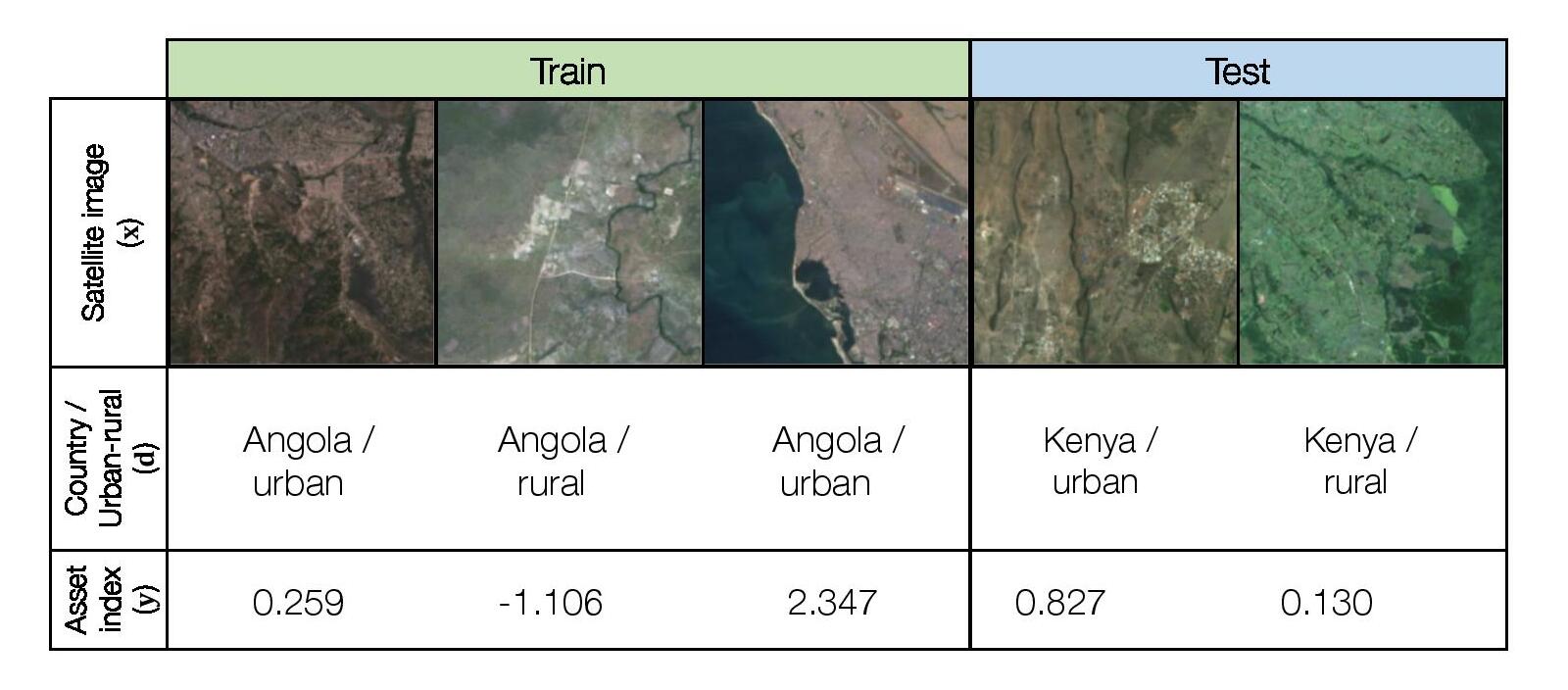
We consider a hybrid domain generalization and subpopulation shift problem, where the input x is a multispectral LandSat satellite image with 8 channels (resized to 224 x 224 pixels), the output y is a real-valued asset wealth index computed from Demographic and Health Surveys (DHS) data, and the domain d represents the country the image was taken in and whether the image is of an urban or rural area. We aim to solve both a domain generalization problem across country borders and improve subpopulation performance across urban and rural areas.
Dataset citation
@article{yeh2020using,
title={Using publicly available satellite imagery and deep learning to understand economic well-being in Africa},
author={Yeh, Christopher and Perez, Anthony and Driscoll, Anne and Azzari, George and Tang, Zhongyi and Lobell, David and Ermon, Stefano and Burke, Marshall},
journal={Nature Communications},
year={2020},
publisher={Nature Publishing Group}
}
License
LandSat/DMSP/VIIRS data is U.S. Public Domain.
Amazon

Motivation
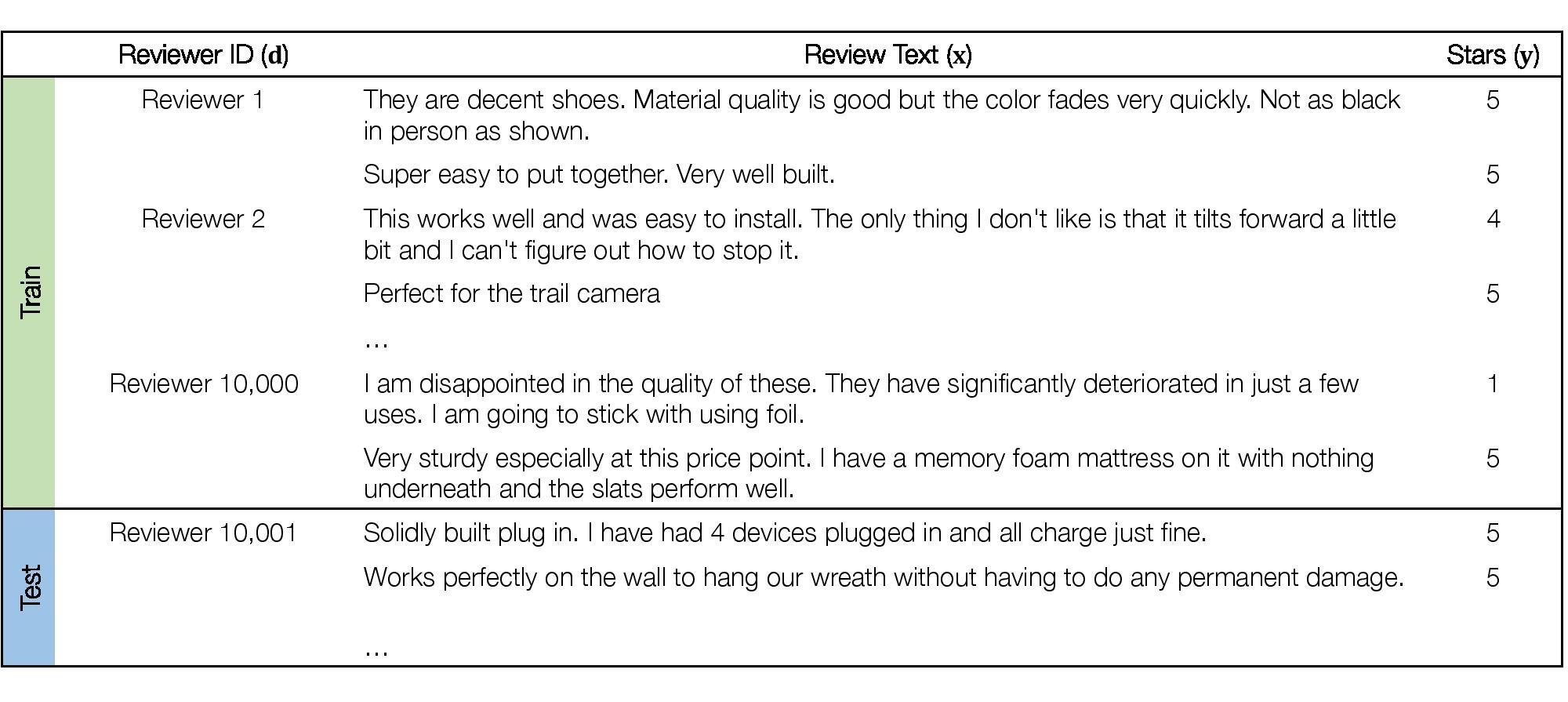
In many consumer-facing ML applications, models are trained on data collected on one set of users and then deployed across a wide range of potentially new users. These models can perform well on average but poorly on some individuals (Li et al., 2019b; Caldas et al., 2018; Geva et al., 2019; Tatman, 2017; Koenecke et al., 2020). These large performance disparities across users are practical concerns in consumer-facing applications, and they can also indicate that models are exploiting biases or spurious correlations in the data (Badgeley et al., 2019; Geva et al., 2019).
We study this issue of inter-individual performance disparities on a variant of the Amazon Reviews dataset (Ni et al., 2019).
Problem setting
We consider a hybrid domain generalization and subpopulation problem where the domains correspond to different reviewers. The task is multi-class sentiment classification, where the input x is the text of a review, the label y is a corresponding star rating from 1 to 5, and the domain d is the identifier of the reviewer who wrote the review. Our goal is to perform consistently well across a wide range of reviewers, i.e., to achieve high tail performance on different subpopulations of reviewers in addition to high average performance. In addition, we consider disjoint set of reviewers between training and test time.
Dataset citation
@inproceedings{ni2019justifying,
title={Justifying recommendations using distantly-labeled reviews and fine-grained aspects},
author={Ni, Jianmo and Li, Jiacheng and McAuley, Julian},
booktitle={Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP)},
year={2019}
}
License
None. The original authors request that the data be used for research purposes only.
Py150
Motivation
Code completion models—autocomplete tools used by programmers to suggest subsequent source code tokens, such as the names of API calls—are commonly used to reduce the effort of software development (Robbes et al., 2008; Bruch et al., 2009; Proksch et al., 2015; Nguyen et al., 2015; Franks et al., 2015). These models are typically trained on data collected from existing codebases but then deployed more generally across other codebases, which may have different distributions of API usages (Nita et al., 2010; Proksch et al., 2016; Allamanis & Brockschmidt, 2017). This shift across codebases can cause substantial performance drops in code completion models. Moreover, prior studies of real-world usage of code completion models have noted that these models can generalize poorly on some important subpopulations of tokens such as method names (Hellendoorn et al., 209).
We study this problem using a variant of the Py150 Dataset, originally developed by Raychev et al., 2016 and adapted to a code completion task by (Lu et al., 2021).
Problem setting
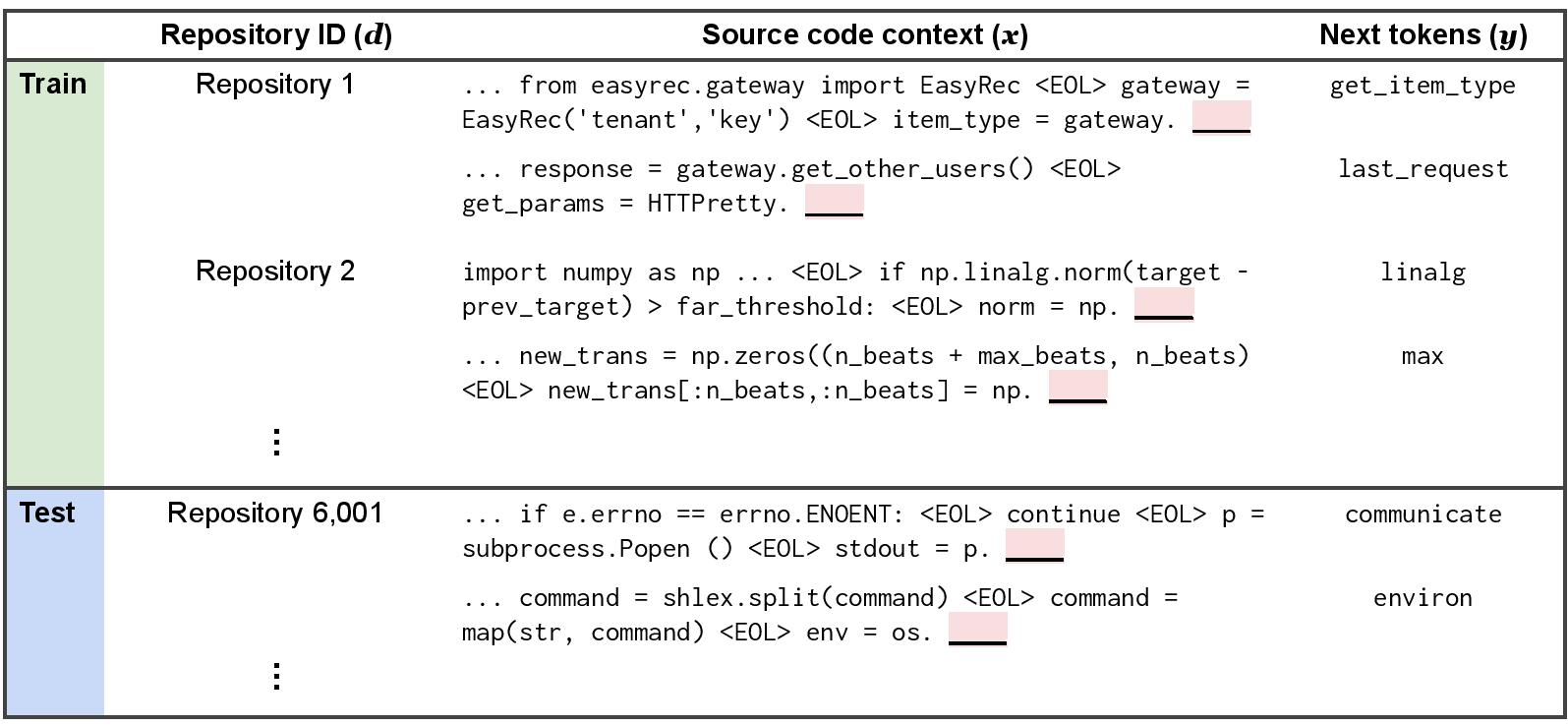
We consider a hybrid domain generalization and subpopulation shift problem, where the domains are codebases (GitHub repositories), and our goal is to learn code completion models that generalize to source code written in new codebases.
Concretely, the input x is a sequence of source code tokens taken from a single file, the label y is the next token (e.g., environ and communicate in the dataset figure), and the domain d is an integer that identifies the repository that the source code belongs to.
We aim to solve both a domain generalization problem across codebases and improve subpopulation performance on class and methods tokens.
Dataset citation
@article{lu2021codexglue,
title={CodeXGLUE: A Machine Learning Benchmark Dataset for Code Understanding and Generation},
author={Lu, Shuai and Guo, Daya and Ren, Shuo and Huang, Junjie and Svyatkovskiy, Alexey and Blanco, Ambrosio and Clement, Colin and Drain, Dawn and Jiang, Daxin and Tang, Duyu and others},
journal={arXiv preprint arXiv:2102.04664},
year={2021}
}
@article{raychev2016probabilistic,
title={Probabilistic model for code with decision trees},
author={Raychev, Veselin and Bielik, Pavol and Vechev, Martin},
journal={ACM SIGPLAN Notices},
year={2016},
}
License
Distributed under the MIT license.